cran_pkgs <- c(
"cutoff", "data.table", "dbscan", "dplyr", "factoextra", "ggrepel", "ggplot2",
"ggside", "htmltools", "htmlwidgets", "irr", "patchwork", "plotly", "reactable",
"reactablefmtr", "scales", "scattermore", "shadowtext", "sparkline", "stringr",
"sp", "UpSetR", "MASS", "e1071", "R.utils", "rlang", "rmarkdown", "uwot",
"future.apply", "progressr", "future", "aricode", "clusterSim", "fpc",
"RColorBrewer", "gtools", "lmerTest", "Matrix", "Seurat", "SeuratObject"
)
cran_missing <- cran_pkgs[!cran_pkgs %in% rownames(installed.packages())]
if (length(cran_missing) > 0) {
install.packages(cran_missing)
}
bioc_pkgs <- c(
"MatrixGenerics", "scater", "scDblFinder",
"SingleCellExperiment", "S4Vectors", "bluster", "dreamlet", "variancePartition"
)
bioc_missing <- bioc_pkgs[!bioc_pkgs %in% rownames(installed.packages())]
if (length(bioc_missing) > 0) {
BiocManager::install(bioc_missing)
}SingleCellMQC v0.7.1
Introduction
SingleCellMQC
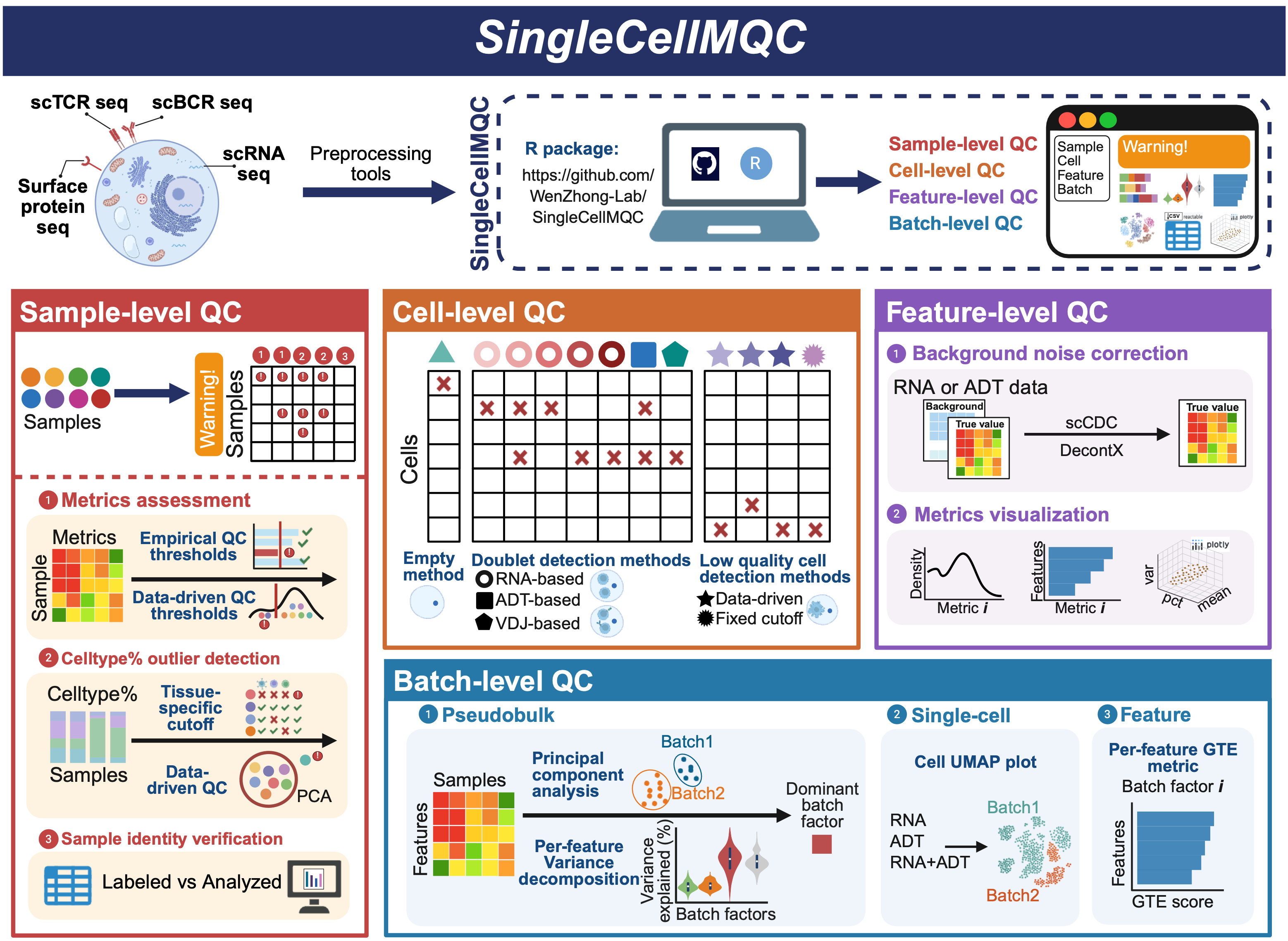
SingleCellMQC: A comprehensive quality control workflow for single-cell single omics or multi-omics. The SingleCellMQC workflow consists of four major QC modules: sample-level QC, cell-level QC, feature-level QC, and batch-level QC. All functions within the SingleCellMQC package are built upon the Seurat and BPCells frameworks and are compatiable with commonly used single-cell analysis pipelines. Each QC module can be executed independently, allowing users to perform QC tasks for specific aims. Moreover, interactive HTML reports summarizing QC results can be generated via the ‘RunReport’ function. In addition, the workflow is optimized for large-scale datasets, and all QC tasks can be executed on a standard laptop without requiring high-performance computing.
R Installation
Before installation, we recommend installing these dependencies first:
You can install the development version of SingleCellMQC :
if (!requireNamespace("devtools", quietly = TRUE)) {
install.packages("devtools")
}
devtools::install_github('WenZhong-Lab/SingleCellMQC')
library(SingleCellMQC)